Objective Tinnitus Detection from fNIRS
Led by Melbourne Bionics Institute, a machine learning workflow for objective tinnitus classification from fNIRS hbo/hbr signals, combining TSFRESH feature engineering, PCA compression, and repeated model evaluation.

Project Summary
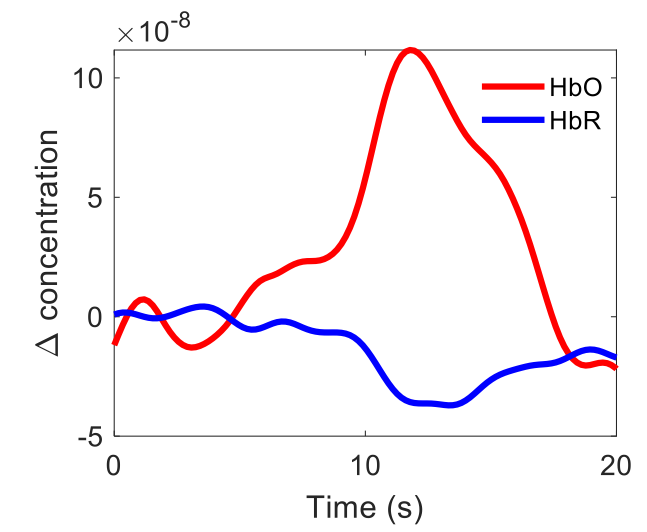
Melbourne Bionics Institute led this project, which investigates whether fNIRS time-series signals can support objective identification of tinnitus cases versus controls. Data includes 32 channels (2 x 16 layout) with both hbo and hbr signals collected across resting and stimulus-triggered sessions.
The pipeline transforms channel-level time-series into rich feature vectors, reduces dimensionality, and benchmarks multiple classifiers over repeated stratified splits using clinical performance metrics.
My Role
I developed and implemented the machine learning modeling workflow with a focus on TSFRESH-based feature engineering for physiological time-series within the Melbourne Bionics Institute-led research program. My contributions included:
- Building data-to-feature pipelines for resting and trigger-mode experiments.
- Designing PCA-based dimensionality reduction for high-dimensional TSFRESH outputs.
- Training and evaluating SVM, Random Forest, and kNN models with repeated validation.
- Producing channel-ranking analyses to identify high-value signal locations.
- Developing a deployable end-to-end GUI that accepts fNIRS sensory records as input and returns ML-based diagnosis output with a severity score and confidence estimate.
Technical Workflow
1) Data Structuring
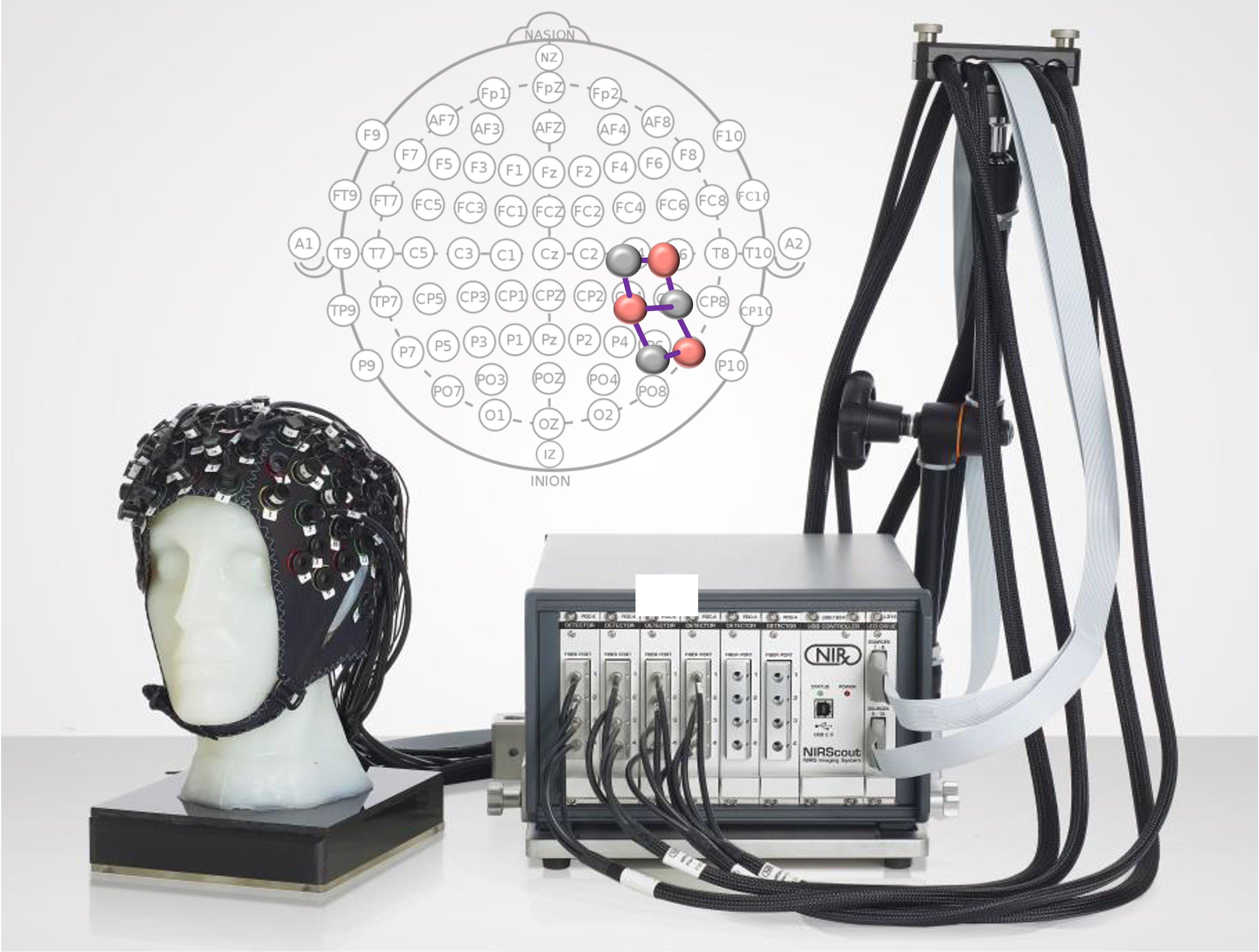

Per-channel hbo/hbr signals are chunked and transformed into modeling tables for subject-level analyses in resting and trigger settings.
2) TSFRESH Features
Automated extraction of comprehensive time-series descriptors is performed with TSFRESH to capture statistical and temporal signal characteristics.
3) Dimensionality Reduction
PCA reduces dimensionality and stabilizes downstream model fitting across channels and trigger conditions.
4) Classification
Repeated train/test evaluation compares SVM, Random Forest, and kNN, reporting AUC, sensitivity, specificity, and accuracy.
5) Channel and ROI Analysis
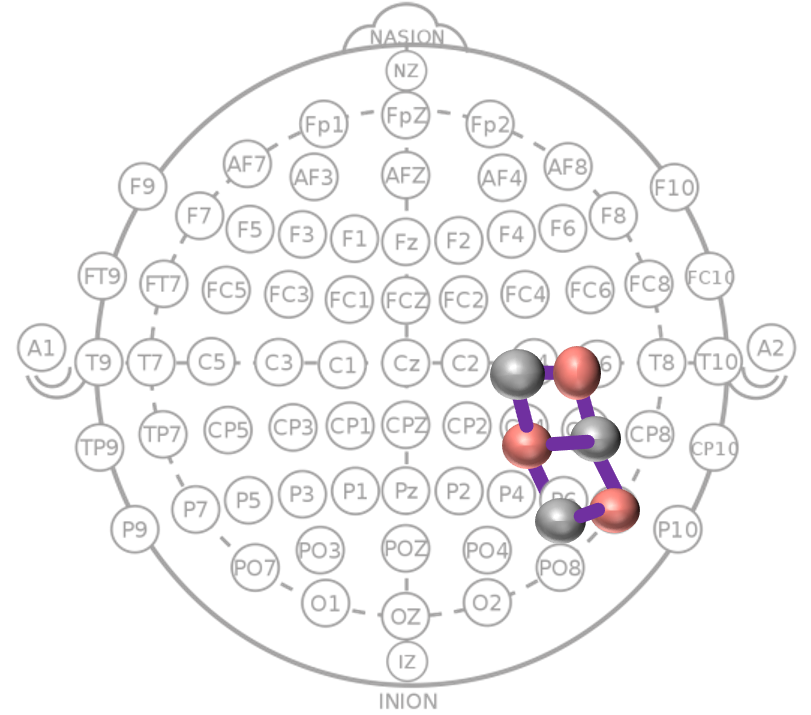
Channel-level and regional analyses (frontal, temporal, occipital) are used to interpret where discriminative signal patterns are strongest.
6) Clinical Translation
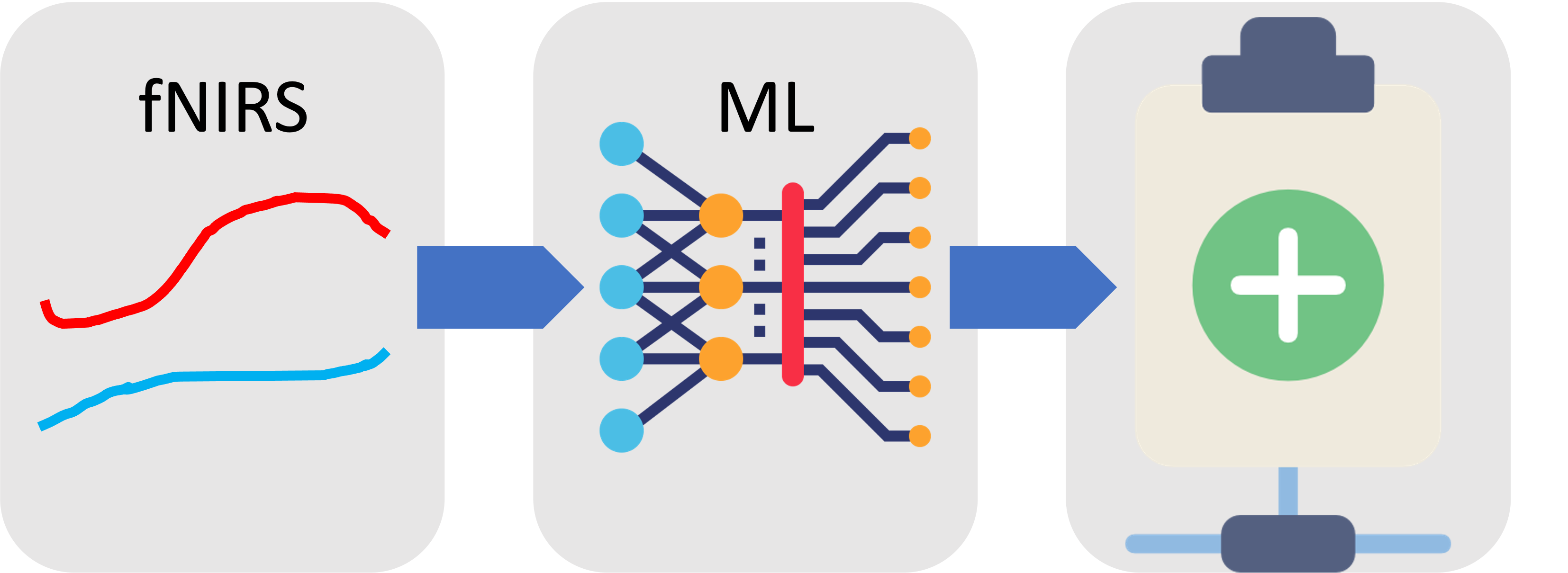
The framework demonstrates how neurophysiological time-series can be converted into reproducible
machine learning evidence for clinical decision-support research.